QMM LIBRARY -- A Mod for LIBLINEAR
A regularized empirical risk minimization software developed at Sabanci University
PROJECT NEWS
- Last updated: Dec.31.2012
- We plan to update our software regularly based on user needs. You can contact Hakan Erdogan for further information.
INTRODUCTION
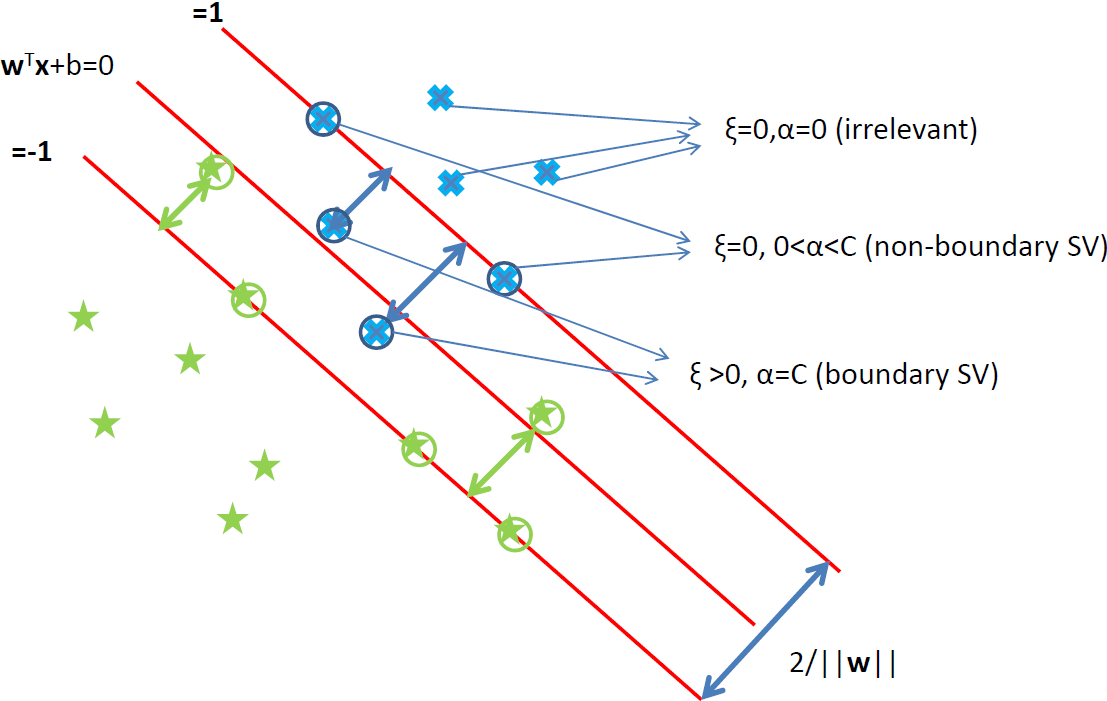
QMM library is a general solver for binary and multi-class linear classifiers optimizing regularized empirical risk.
- Based on majorize minimize (MM) method
- Monotonicity guarantees
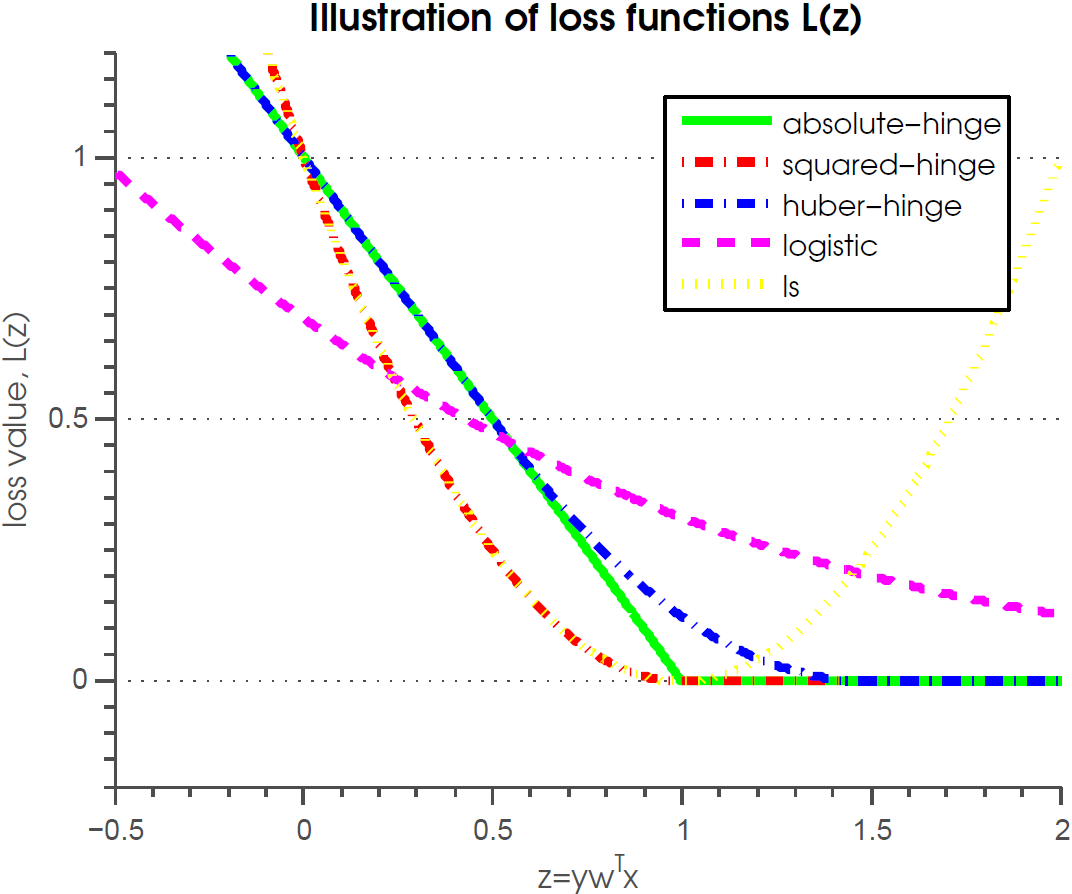
- Possible to use many different loss functions (squared-hinge, Huber-hinge, logistic loss)
- Supports L1, L2 or elastic net regularization (sparse parameter training with L1 regularization)
CONTRIBUTORS (programmers):
(1) I. Saygin Topkaya (2) M. Umut Sen (3) Hakan Erdogan
DOWNLOAD
The software is hosted on github. Visit the github page here for the latest version.
You can directly download the ZIP file or download the tarball as well.
INSTALLATION (on linux):
Windows executables are included under the windows folder.
For linux:
- Run make in the main folder.
- Run make.m from within matlab in the matlab folder.
SIMPLE USE:
- download database files a1a.train and a1a.test from libsvm databases page
- run this command in the command prompt:
- train -B 1 -h 0 -s 50 -u 1 -l 2 -r 1 a1a.train trained_model.txt
- this will generate a file called trained_model.txt which contains a logistic regression binary linear classifier trained using the MMCD method with the optimal curvature
- to test/predict with the trained model file run:
- predict a1a.test trained_model.txt output_file.txt
- for more details, see the argument help below and for even more details read the documentation
PROGRAM ARGUMENTS
QMM library is a modification of the liblinear software. You can use the same input and output formats. We have added new types of solvers to the liblinear software to enable use of our algorithms.
To enable using the QMM library use the options -s 50-55.
Here is a list of arguments for the modified LIBLINEAR train program.
Usage: train [options] training_set_file [model_file]
options:
-s type : set type of solver (default 1)
0 -- L2-regularized logistic regression (primal)
1 -- L2-regularized L2-loss support vector classification (dual)
2 -- L2-regularized L2-loss support vector classification (primal)
3 -- L2-regularized L1-loss support vector classification (dual)
4 -- multi-class support vector classification by Crammer and Singer
5 -- L1-regularized L2-loss support vector classification
6 -- L1-regularized logistic regression
7 -- L2-regularized logistic regression (dual)
11 -- L2-regularized L2-loss epsilon support vector regression (primal)
12 -- L2-regularized L2-loss epsilon support vector regression (dual)
13 -- L2-regularized L1-loss epsilon support vector regression (dual)
50 -- MMCD - please specify loss, curv
51 -- MMCD_SM - soft-max method
52 -- MMCD_SG - sub-gradient
53 -- MMCD_SIMPLE - please specify loss, curv
54 -- MMGCD - please specify loss, curv
55 -- MMCG - please specify loss, curv
-l loss_type : L1, L2, LOG, HU1, HU2, LS
0 -- L1
1 -- L2
2 -- LOG
3 -- HU1
4 -- HU2
5 -- LS
-u curv_type : MC, OC, NC
0 -- MC
1 -- OC
2 -- NC
for -s 50, 54, and 55 :
3 -- Start with MC and continue with NC after 1st iteration
4 -- Start with OC and continue with NC after 1st iteration
-r alpha : regularization parameter, between 0 and 1
0 -- L1 regularization
1 -- L2 regularization
0 < r <1 -- elastic net
-t tau : loss parameter for HU1, HU2 and L1 losses, default 0.5
-n epsi : curv parameter, minimum curvature for NC, typical 0.001
-i initmodel : initial model file to start iterations
-g cg_tol : conjugate gradient tolerance for MMCG, default 0.001
-d cd_tol : coordinate decent tolerance for MMCD, default 0.01
-f n : only for MMCD, reset active coordinates every nth iteration, default 10
-h chat_level : how much should I talk?
0 -- minimal
1 -- calc and print obj
for MMCD_SIMPLE, MMGCD and MMCG:
1 - running time
2 - |f'(w)|_2 and |f'(w)|_inf
3 - objective value
4 - training and testing accuracy; mention test file with -x
-x filename : test file to be used if chat_level>=4
-c cost : set the parameter C (default 1)
-p epsilon : set the epsilon in loss function of epsilon-SVR (default 0.1)
-e epsilon : set tolerance of termination criterion
-s 0 and 2
|f'(w)|_2 <= eps*min(pos,neg)/l*|f'(w0)|_2,
where f is the primal function and pos/neg are # of
positive/negative data (default 0.01)
-s 11
|f'(w)|_2 <= eps*|f'(w0)|_2 (default 0.001)
-s 1, 3, 4, and 7
Dual maximal violation <= eps; similar to libsvm (default 0.1)
-s 5 and 6
|f'(w)|_1 <= eps*min(pos,neg)/l*|f'(w0)|_1,
where f is the primal function (default 0.01)
-s 12 and 13
|f'(alpha)|_1 <= eps |f'(alpha0)|,
where f is the dual function (default 0.1)
-s 50, 54, and 55
|w-w^prev|_inf <= eps*|w|_inf,
-S structure : trains structured weights;
structure is s for symmetric, a for antisymmetric
or a filename of the free transform matrix (default none)
-B bias : if bias >= 0, instance x becomes [x; bias]; if < 0, no bias term added (default -1)
-wi weight: weights adjust the parameter C of different classes (see README for details)
-v n: n-fold cross validation mode
-q : quiet mode (no outputs)
DOCUMENTATION AND CODES
The codes can be received from the github page.
For technical description, see our technical report (coming soon).
For code documentation, you can see the doxygen documentation page (the doc folder) that is in the github package.Or you can visit this link for doxygen source code documentation.
For abstract of the project in which this software was built English / Turkish
PAPERS RELATED TO OUR WORK
-
Mehmet Umut Şen, Hakan Erdoğan, "Linear classifier combination and selection using group sparse regularization and hinge loss," Pattern Recognition Letters, vol. 34, no. 3, pp. 265-274, Feb. 2003.
-
Mehmet Umut Sen, Hakan Erdogan, "Basit Birlestirici Tipleri için Dogrusal Olmayan Siniflandirici Birlestirme," IEEE SIU 2011, Kemer, Nisan 2011. (in Turkish)
-
Ibrahim Saygin Topkaya, Mehmet Umut Sen, Mustafa Berkay Yilmaz, Hakan Erdogan, "Görsel-Isitsel Tandem Siniflandiricilar ve Birlesimleri ile Konusma Tanima Basarisini Arttirma," IEEE SIU 2011, Kemer, Nisan 2011. (in Turkish)
Please
send comments and suggestions to Hakan Erdogan.